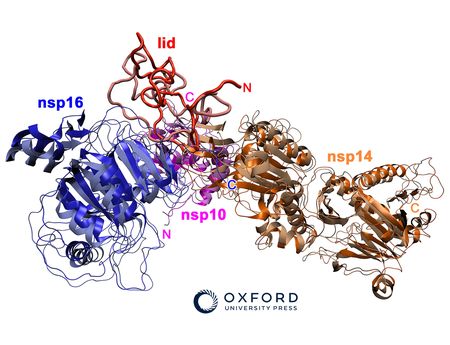
We study different translation control mechanisms, which regulate the production of specific sets of proteins by chemical modifications of tRNA molecules. Every protein in the cell is produced by the ribosome, which uses transfer RNA (tRNA) molecules to translate the sequence information coded in mRNAs into correctly assembled poly-peptide chains. The decoding/translation of genetic information is based on the recognition of a respective codon by its corresponding tRNA anticodon triplet.
The lab is focusing on understanding the molecular mechanisms that lead to the specific base modifications in anticodons of tRNAs. These modifications have a strong influence on the efficiency and accuracy of the codon-anticodon pairing and therefore regulate the translational rates and folding dynamics of protein synthesis. Recent findings have shown that alterations of these modification pathways play important roles in the onset of certain neurodegenerative diseases and cancer.
https://www.uj.edu.pl/pl_PL/web/malopolskie-centrum-biotechnologii/glatt-lab-max-planck
ATMP products
Now we are ready to start produce HE-ATMP with cooperation: with clinical units for the use of ATMP products in treatment with biotechnological companies which strive for implementation of medicinal products Also realization of scientific projects as a part of which medical experiments with the use of ATMP are anticipated – SUBCONTRACTING OF PROJECTS WHICH ASSUME THE USE OF ATMP- will be possible. Particularly, in relation to the best experience of the qualified bank staff, realization of projects related to the use of skin cells in treatment of burns, chronical wounds and pathologic skin lesions, is assumed. Now, as one of the few Cell Banks in Poland, we have the necessary authorization to manufacture several HE - ATMP products as well as years of experience which allow us to effectively implement the intended assumptions of the project. The Bank head will again apply for funding of the project related to implementation of 3 ATMP products: Clinical application of keratinocytes in treating burns and wounds which are hard to cure Mesenchymal adipose-derived stem cells (ASC): treating soft tissue defects of the rectum area The use of autologous chondrocytes in treating cartilaginous tissue defects We plan to start cooperation with two companies, for which ATMP products will be produced.
 Web Content Display
Web Content Display
 Web Content Display
Web Content Display